Entangled networks#
For these examples, we use the models of a network liquid from Neophytou et al., PNAS, 121, e2406890121 which is known to densify via two successive liquid-liquid phase transitions.
The system of interest is a coarse-grained model of DNA dendrimers, where each dendrimer is represented by a repulsive spherical core decorated with four attractive patches in a tetrahedral arrangement.
At low pressures, an unentangled low-density liquid (LDL) forms.
At intermediate pressures, an entangled high-density liquid (HDL) forms.
At high pressures, an entangled very high-density liquid (vHDL) forms.
[1]:
import math
import random
from collections import defaultdict
from itertools import combinations
from typing import Any, Iterable
import matplotlib.pyplot as plt
import numpy as np
from tqdm import tqdm
import topo_metrics as tm
import topo_metrics.analysis as tm_ana
from topo_metrics._julia_wrappers import run_bond_distance_rdf
%config InlineBackend.figure_format = 'svg'
radial distribution functions by bond separation#
The first measure used to identify the onset of entanglement is to decompose the radial distribution function (RDF) into contributions derived from particles separated by \(D\) bonds. This allows for assessing whether particles close in space are separated by multiple bonds in the network.
[2]:
all_rs = []
all_gD = []
for index in range(1, 101):
topo = tm.Topology.from_conflink("data/conflink_p2.0.dat", index=-index)
assert topo.lattice is not None, "Topology must have lattice information."
rs, gD, _ = run_bond_distance_rdf(
positions=topo.cartesian_coordinates,
edges=topo.edges,
cell_lengths=topo.lattice.lengths,
cell_angles=topo.lattice.angles,
dmax=6,
rmax=50.0,
dr=0.1,
)
all_rs.append(rs)
all_gD.append(gD)
[3]:
for rs in all_rs[1:]:
assert np.allclose(rs, all_rs[0])
rs = all_rs[0]
std_gD = np.std(np.array(all_gD), axis=0)
mean_gD = np.mean(np.array(all_gD), axis=0)
fig, ax = plt.subplots()
for D in [1, 2, 3, 4, 5, 6]:
ax.plot(rs / 8, mean_gD[D - 1], label=f"D={D}", alpha=0.9)
if D in [1, 2]:
ax.fill_between(rs / 8, 0, mean_gD[D - 1], alpha=0.1)
ax.set_xlabel("r")
ax.set_ylabel("g(r)")
ax.set_ylim(-0.05, 1.25)
ax.legend(ncol=2, frameon=False)
plt.show()
[ ]:
all_rs = []
all_gD = []
for index in range(1, 101):
topo = tm.Topology.from_conflink("data/conflink_p20.0.dat", index=-index)
assert topo.lattice is not None, "Topology must have lattice information."
rs, gD, _ = run_bond_distance_rdf(
positions=topo.cartesian_coordinates,
edges=topo.edges,
cell_lengths=topo.lattice.lengths,
cell_angles=topo.lattice.angles,
dmax=6,
rmax=50.0,
dr=0.1,
)
all_rs.append(rs)
all_gD.append(gD)
for rs in all_rs[1:]:
assert np.allclose(rs, all_rs[0])
rs = all_rs[0]
std_gD = np.std(np.array(all_gD), axis=0)
mean_gD = np.mean(np.array(all_gD), axis=0)
fig, ax = plt.subplots()
for D in [1, 2, 3, 4, 5, 6]:
ax.plot(rs / 8, mean_gD[D - 1], label=f"D={D}", alpha=0.9)
if D in [1, 2]:
ax.fill_between(rs / 8, 0, mean_gD[D - 1], alpha=0.1)
ax.set_xlabel("r")
ax.set_ylabel("g(r)")
ax.set_ylim(-0.05, 1.25)
ax.legend(ncol=2, frameon=False)
plt.show()
[ ]:
all_rs = []
all_gD = []
for index in range(1, 101):
topo = tm.Topology.from_conflink("data/conflink_p32.0.dat", index=-index)
assert topo.lattice is not None, "Topology must have lattice information."
rs, gD, _ = run_bond_distance_rdf(
positions=topo.cartesian_coordinates,
edges=topo.edges,
cell_lengths=topo.lattice.lengths,
cell_angles=topo.lattice.angles,
dmax=6,
rmax=50.0,
dr=0.1,
)
all_rs.append(rs)
all_gD.append(gD)
for rs in all_rs[1:]:
assert np.allclose(rs, all_rs[0])
rs = all_rs[0]
std_gD = np.std(np.array(all_gD), axis=0)
mean_gD = np.mean(np.array(all_gD), axis=0)
fig, ax = plt.subplots()
for D in [1, 2, 3, 4, 5, 6]:
ax.plot(rs / 8, mean_gD[D - 1], label=f"D={D}", alpha=0.9)
if D in [1, 2]:
ax.fill_between(rs / 8, 0, mean_gD[D - 1], alpha=0.1)
ax.set_xlabel("r")
ax.set_ylabel("g(r)")
ax.set_ylim(-0.05, 1.25)
ax.legend(ncol=2, frameon=False)
plt.show()
coiled rings#
Coiled rings are cyclic paths in the network whose geometry is strongly distorted away from a planar-like conformation. In this sense, coiled rings exhibit supercoiling in three dimensions, i.e., the closed path winds around itself.
The degree of coiling (and, more generally, self-entanglement of the ring backbone) is quantified by the writhe of the cyclic path \(C_i\),
Here, \(\mathbf r_i\) and \(\mathbf r_i'\) denote two points on the same closed curve \(C_i\), and \(d\mathbf r_i\), \(d\mathbf r_i'\) are the corresponding infinitesimal tangent vectors along the curve.
Geometrically, \(Wr_i\) measures how many times the closed path loops around itself in 3D. Approximately planar rings have \(Wr_i \approx 0\), while strongly coiled rings have \(|Wr_i|\gtrsim 1\) (with larger \(|Wr_i|\) indicating more pronounced self-winding). When chirality is not of interest, one often reports \(|Wr_i|\), since the sign of \(Wr_i\) encodes handedness.
[6]:
topo = tm.Topology.from_conflink("data/conflink_p2.0.dat")
rings = topo.get_rings(depth=8)
topo
[6]:
Topology(nodes=1000, edges=3982, has_lattice=True)
[7]:
all_writhes = defaultdict(list)
for ring in rings:
if len(ring) > 10:
continue
writhe, _ = ring.writhe_and_acn() # type: ignore
all_writhes[len(ring)].append(abs(writhe))
bins = np.linspace(0, 1.25, 50)
fig, ax = tm_ana.plot_writhe_distributions(all_writhes, bins=bins)
ax.set_xlim(0, 1.25)
plt.show()
[8]:
topo = tm.Topology.from_conflink("data/conflink_p20.0.dat")
rings = topo.get_rings(depth=8)
all_writhes = defaultdict(list)
for ring in rings:
if len(ring) > 10:
continue
writhe, _ = ring.writhe_and_acn() # type: ignore
all_writhes[len(ring)].append(abs(writhe))
fig, ax = tm_ana.plot_writhe_distributions(all_writhes, bins=bins)
ax.set_xlim(0, 1.25)
plt.show()
[9]:
topo = tm.Topology.from_conflink("data/conflink_p32.0.dat")
rings = topo.get_rings(depth=8)
all_writhes = defaultdict(list)
for ring in rings:
if len(ring) > 10:
continue
writhe, _ = ring.writhe_and_acn() # type: ignore
all_writhes[len(ring)].append(abs(writhe))
fig, ax = tm_ana.plot_writhe_distributions(all_writhes, bins=bins)
ax.set_xlim(0, 1.25)
plt.show()
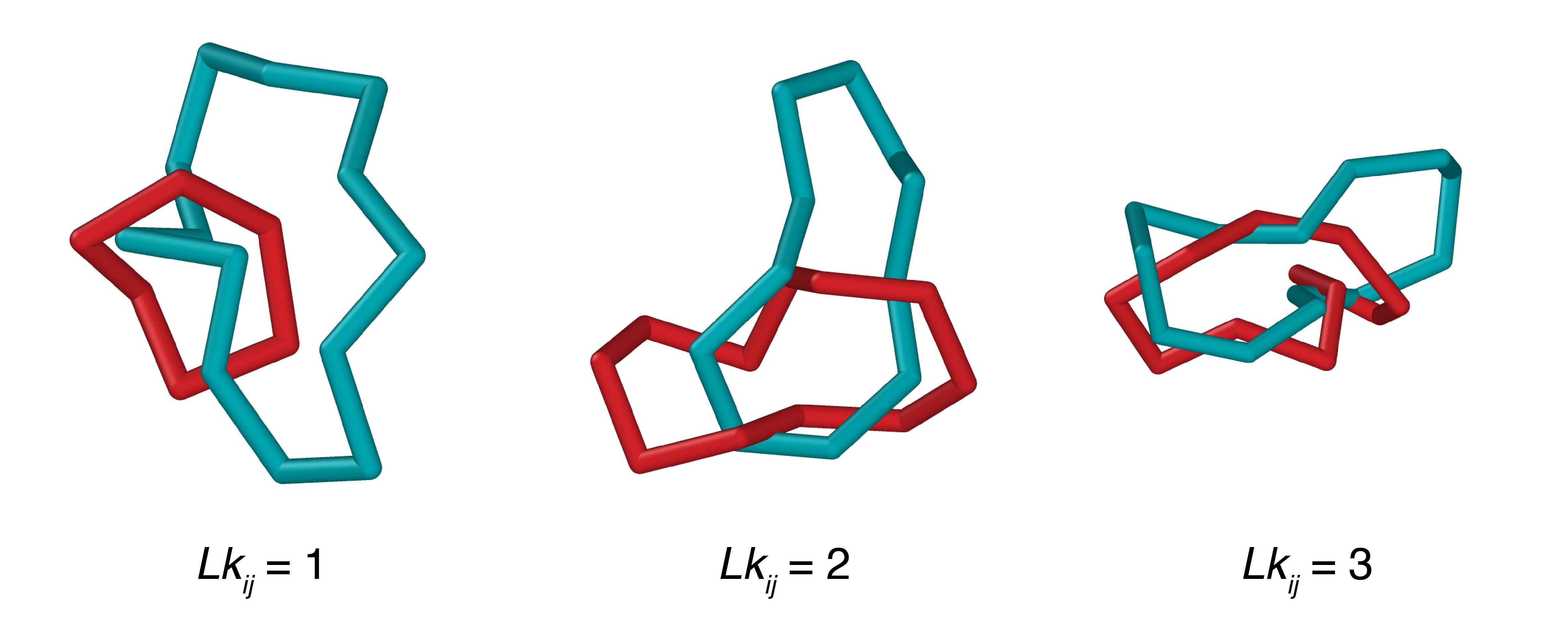
linked rings#
Linked rings are two (or more) disjoint rings (i.e., rings that do not share any vertices) which are entangled with one another in such a way that they cannot be unlinked without at least one bond being broken.
[10]:
from topo_metrics import Node
from topo_metrics.io.lmp import write_rings_lammps_full
from topo_metrics.knots import linking_number_method_1a
from topo_metrics.ring_geometry import RingGeometry
def circle_xy(n=400, r=1.0, center=(0.0, 0.0, 0.0), z=0.0) -> RingGeometry:
"""circle in plane z = center[2] + z."""
t = np.linspace(0.0, 2.0 * math.pi, n, endpoint=False)
x = center[0] + r * np.cos(t)
y = center[1] + r * np.sin(t)
zz = np.full_like(t, center[2] + z)
coords = np.column_stack([x, y, zz])
return RingGeometry(
tuple(
Node(node_id=node_id, cart_coord=pos)
for node_id, pos in enumerate(coords, start=1)
)
)
def circle_xz(n=400, r=1.0, center=(1.0, 0.0, 0.0)):
"""circle in plane y = center[1]"""
t = np.linspace(0.0, 2.0 * math.pi, n, endpoint=False)
x = center[0] + r * np.cos(t)
y = np.full_like(t, center[1])
z = center[2] + r * np.sin(t)
coords = np.column_stack([x, y, z])
return RingGeometry(
tuple(
Node(node_id=node_id, cart_coord=pos)
for node_id, pos in enumerate(coords, start=1)
)
)
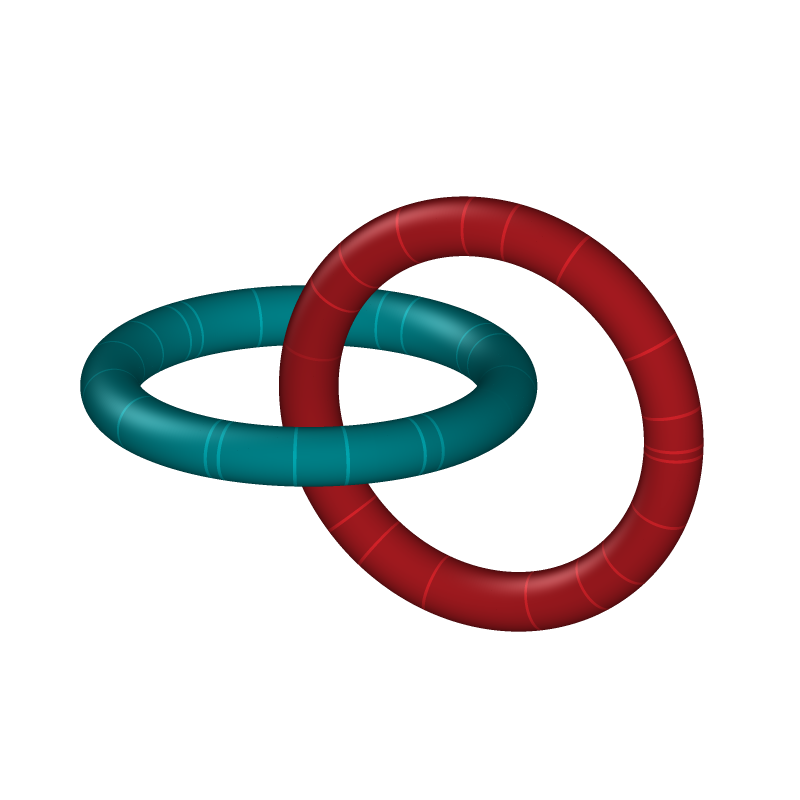
Hopf link#
The Hopf link is the simplest nontrivial link with two components. It consists of two closed loops linked exactly once, meaning the two components cannot be separated without cutting. In an oriented setting, its linking number is \(\mathrm{Lk} = \pm 1\) (the sign depends on the chosen orientations). The Hopf link is named after Heinz Hopf.

[ ]:
ring1 = circle_xy(n=400, r=1.0)
ring2 = circle_xz(n=400, r=1.0, center=(1, 0, 0))
write_rings_lammps_full(
rings=[ring1, ring2],
filepath="configs/hopf.data",
cell=np.diag([10.0, 10.0, 10.0]),
pbc=True,
)
lk = linking_number_method_1a(ring1.positions, ring2.positions)
print(f"Lk (float): {lk:.0f}")
Lk (float): -1
To illustrate how the linking number depends on orientation, we compute it for the original ordering of one ring and again after reversing that ring’s vertex order.
[12]:
lk1 = linking_number_method_1a(ring1.positions, ring2.positions)
lk2 = linking_number_method_1a(ring1.positions, ring2.positions[::-1])
print("Lk forward:", int(round(lk1)), "\nLk reversed:", int(round(lk2)))
Lk forward: -1
Lk reversed: 1
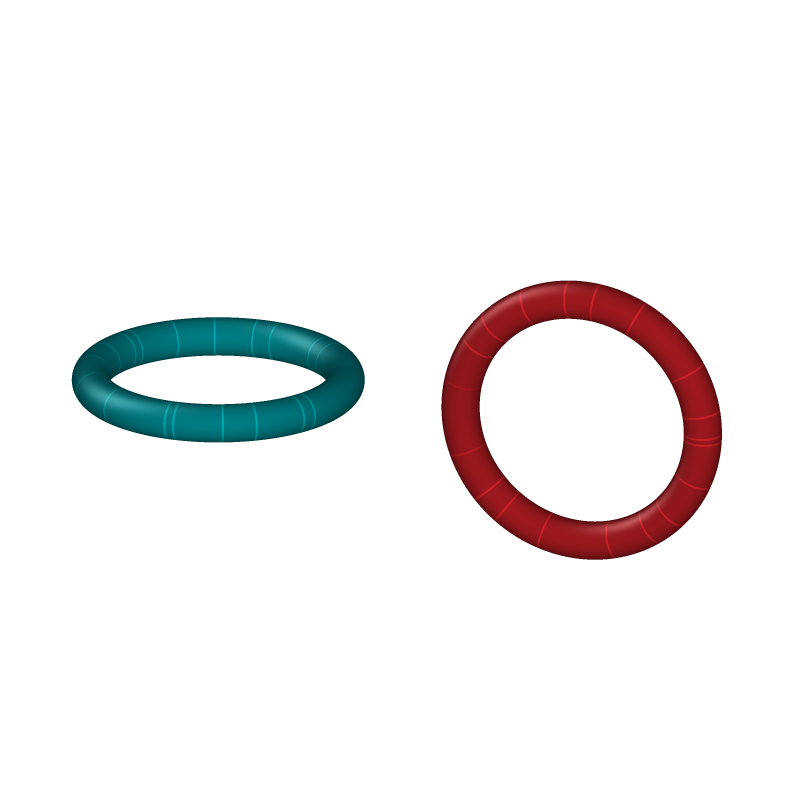
As a sanity check, we place the second ring far from the first (so the two components are unlinked) and verify that the computed linking number is zero.

[13]:
ring2_far = circle_xz(n=400, r=1.0, center=(3, 0, 0))
write_rings_lammps_full(
rings=[ring1, ring2_far],
filepath="configs/separate.data",
cell=np.diag([10.0, 10.0, 10.0]),
pbc=True,
)
lk = linking_number_method_1a(ring1.positions, ring2_far.positions)
print("Lk:", int(round(lk)))
Lk: 0
Now we apply the same ring-topology workflow—ring extraction from the bonded tetrahedral network followed by pairwise linking-number evaluation—to the vHDA tetrahedral network models explored earlier. In particular, we focus on comparing the lowest and highest state points. This analysis is closely motivated by the framework of Neophytou et al. (PNAS 121, e2406890121), who argue that densification in tetrahedral network liquids is driven by the emergence of linked rings and proceeds via a hierarchy of topological transitions.
[14]:
topo_p2 = tm.Topology.from_conflink("data/conflink_p2.0.dat")
rings_p2 = topo_p2.get_rings(depth=8)
assert topo_p2.lattice is not None, "Topology must have lattice information."
n_rings = len(rings_p2)
n_pairs = n_rings * (n_rings - 1) // 2
linked_rings_p2 = []
for ringA, ringB in tqdm(combinations(rings_p2, 2), total=n_pairs):
lk, img = ringA.linking_number(ringB, cell=topo_p2.lattice.matrix)
lk_rounded = np.round(abs(lk), 8)
linked_rings_p2.append((ringA, ringB, lk, img))
100%|██████████| 1865346/1865346 [06:48<00:00, 4562.66it/s]
[15]:
topo_p32 = tm.Topology.from_conflink("data/conflink_p32.0.dat")
rings_p32 = topo_p32.get_rings(depth=8)
assert topo_p32.lattice is not None, "Topology must have lattice information."
n_rings = len(rings_p32)
n_pairs = n_rings * (n_rings - 1) // 2
linked_rings_p32 = []
for ringA, ringB in tqdm(combinations(rings_p32, 2), total=n_pairs):
lk, img = ringA.linking_number(ringB, cell=topo_p32.lattice.matrix)
lk_rounded = np.round(abs(lk), 8)
linked_rings_p32.append((ringA, ringB, lk, img))
100%|██████████| 13889085/13889085 [1:48:27<00:00, 2134.35it/s]
[ ]:
def lk_int_counts(linked_rings) -> tuple[np.ndarray, np.ndarray]:
"""Get counts of integer linking numbers from linked_rings list."""
lk = np.array([abs(x[2]) for x in linked_rings], dtype=float)
lk_i = np.rint(lk).astype(int)
vals, counts = np.unique(lk_i, return_counts=True)
return vals, counts
vals_p2, counts_p2 = lk_int_counts(linked_rings_p2)
vals_p32, counts_p32 = lk_int_counts(linked_rings_p32)
vals = np.union1d(vals_p2, vals_p32)
c2 = np.zeros_like(vals, dtype=int)
c32 = np.zeros_like(vals, dtype=int)
map_p2 = dict(zip(vals_p2, counts_p2))
map_p32 = dict(zip(vals_p32, counts_p32))
for i, v in enumerate(vals):
c2[i] = map_p2.get(v, 0)
c32[i] = map_p32.get(v, 0)
fig, ax = plt.subplots()
w = 0.38
x = np.arange(len(vals))
ax.bar(
x - w / 2,
c2,
width=w,
color="#00A6AF",
alpha=0.9,
label="low pressure (p=2)",
)
ax.bar(
x + w / 2,
c32,
width=w,
color="#D22128",
alpha=0.9,
label="high pressure (p=32)",
)
ax.set_xticks(x)
ax.set_xticklabels(vals)
ax.set_xlabel("|Lk|")
ax.set_yscale("log")
ax.set_ylabel("counts")
ax.legend(frameon=False)
for xi, c in zip(x - w / 2, c2):
if c > 0:
ax.text(xi, c, f"{c}", ha="center", va="bottom")
for xi, c in zip(x + w / 2, c32):
if c > 0:
ax.text(xi, c, f"{c}", ha="center", va="bottom")
plt.show()
[ ]:
def random_linking_example(
linked_rings: Iterable[tuple[Any, Any, float, Any]],
lk_int: int,
*,
tol: float = 1e-6,
rng: random.Random | None = None,
) -> tuple[Any, Any, float, Any]:
"""
Return a random (ring1, ring2, lk_val, img_val) whose lk_val is the
specified integer lk_int.
"""
if rng is None:
rng = random.Random()
matches = []
target = float(lk_int)
for ring1, ring2, lk_val, img_val in linked_rings:
if abs(float(lk_val) - target) <= tol:
matches.append((ring1, ring2, lk_val, img_val))
if not matches:
raise ValueError(
f"No examples found with linking number {lk_int} within tol={tol}"
)
return rng.choice(matches)
[18]:
assert topo_p32.lattice is not None, "Topology must have lattice information."
for lk_target in (1, 2, 3):
ring1, ring2, lk_val, img_val = random_linking_example(
linked_rings=linked_rings_p32, lk_int=lk_target
)
packed = write_rings_lammps_full(
rings=[ring1, ring2],
filepath=f"configs/lk_{lk_target}.data",
cell=topo_p32.lattice.matrix,
pbc=True,
images=np.array([[0, 0, 0], img_val], dtype=int),
)
[ ]: